| << Chapter < Page | Chapter >> Page > |
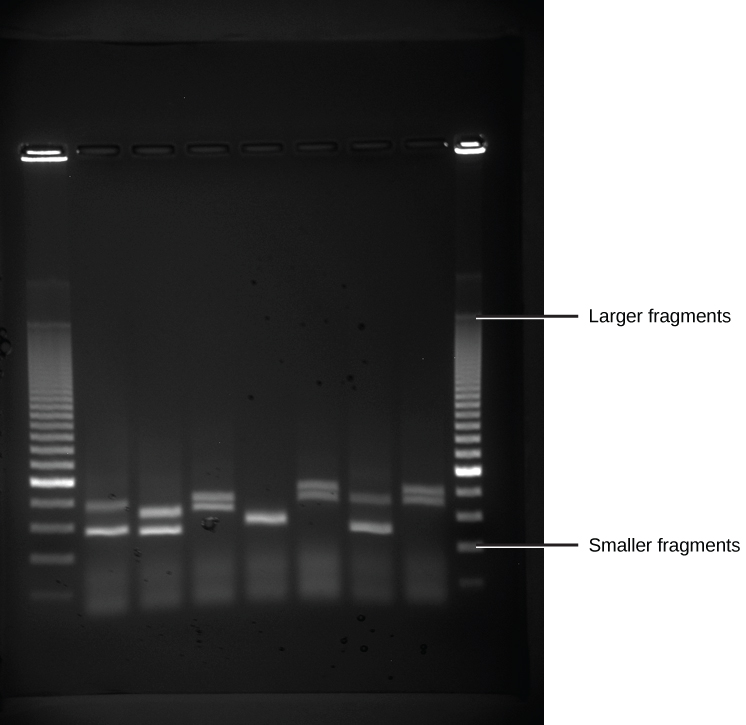
Because nucleic acids are negatively charged ions at neutral or alkaline pH in an aqueous environment, they can be moved by an electric field. Gel electrophoresis is a technique used to separate charged molecules on the basis of size and charge. The nucleic acids can be separated as whole chromosomes or as fragments. The nucleic acids are loaded into a slot at one end of a gel matrix, an electric current is applied, and negatively charged molecules are pulled toward the opposite end of the gel (the end with the positive electrode). Smaller molecules move through the pores in the gel faster than larger molecules; this difference in the rate of migration separates the fragments on the basis of size. The nucleic acids in a gel matrix are invisible until they are stained with a compound that allows them to be seen, such as a dye. Distinct fragments of nucleic acids appear as bands at specific distances from the top of the gel (the negative electrode end) that are based on their size ( [link] ). A mixture of many fragments of varying sizes appear as a long smear, whereas uncut genomic DNA is usually too large to run through the gel and forms a single large band at the top of the gel.
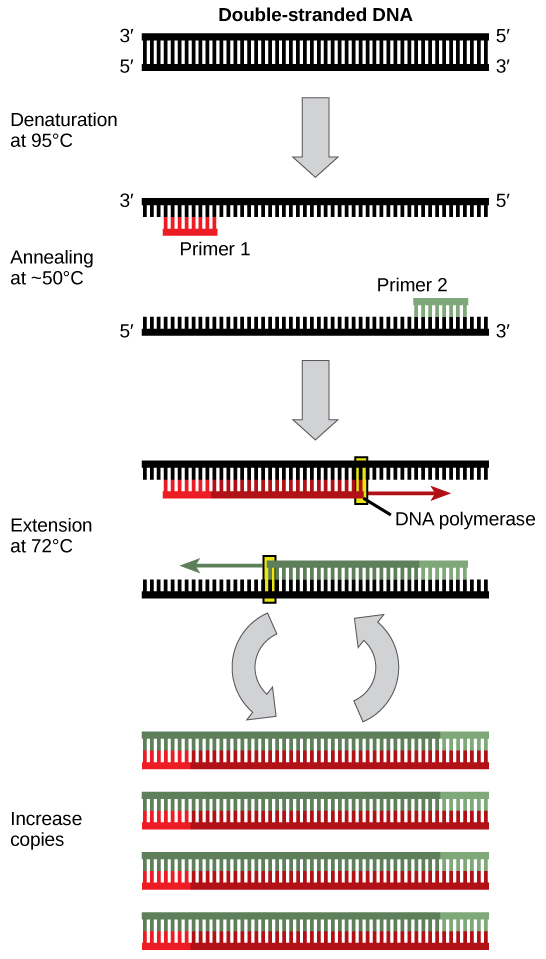
DNA analysis often requires focusing on one or more specific regions of the genome. It also frequently involves situations in which only one or a few copies of a DNA molecule are available for further analysis. These amounts are insufficient for most procedures, such as gel electrophoresis. Polymerase chain reaction (PCR) is a technique used to rapidly increase the number of copies of specific regions of DNA for further analyses ( [link] ). PCR uses a special form of DNA polymerase, the enzyme that replicates DNA, and other short nucleotide sequences called primers that base pair to a specific portion of the DNA being replicated. PCR is used for many purposes in laboratories. These include: 1) the identification of the owner of a DNA sample left at a crime scene; 2) paternity analysis; 3) the comparison of small amounts of ancient DNA with modern organisms; and 4) determining the sequence of nucleotides in a specific region.
In general, cloning means the creation of a perfect replica. Typically, the word is used to describe the creation of a genetically identical copy. In biology, the re-creation of a whole organism is referred to as “reproductive cloning.” Long before attempts were made to clone an entire organism, researchers learned how to copy short stretches of DNA—a process that is referred to as molecular cloning.
Cloning allows for the creation of multiple copies of genes, expression of genes, and study of specific genes. To get the DNA fragment into a bacterial cell in a form that will be copied or expressed, the fragment is first inserted into a plasmid. A plasmid (also called a vector in this context) is a small circular DNA molecule that replicates independently of the chromosomal DNA in bacteria. In cloning, the plasmid molecules can be used to provide a "vehicle" in which to insert a desired DNA fragment. Modified plasmids are usually reintroduced into a bacterial host for replication. As the bacteria divide, they copy their own DNA (including the plasmids). The inserted DNA fragment is copied along with the rest of the bacterial DNA. In a bacterial cell, the fragment of DNA from the human genome (or another organism that is being studied) is referred to as foreign DNA to differentiate it from the DNA of the bacterium (the host DNA).

Notification Switch
Would you like to follow the 'University of georgia concepts of biology' conversation and receive update notifications?